Gene PhylogenyWhat is phylogeny?
Phylogeny is a study of evolution that focuses on the relatedness of species based on a common ancestor. To understand relatedness between species, nucleotide sequence comparisons can provide insight into the molecular alterations that have accumulated within species and are used to identify changes that occurred after the divergence of the species. To visualize the cumulative variation among organisms, phylogentic trees can be constructed, illustrating the relatedness among species or individual genes and the divergence of species throughout time.
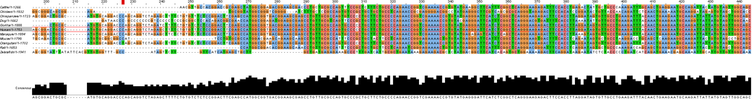
Sequence AlignmentTo begin the process of creating phylogenetic trees, the sequences of the genes of each organism to be studied must be aligned and scored to determine the degree to which the sequences are identical. Clustal Omega is an online multiple sequence alignment program that provides a range of outputs to analyze sequence alignments and was used to align and score the FANCL homologs [1].
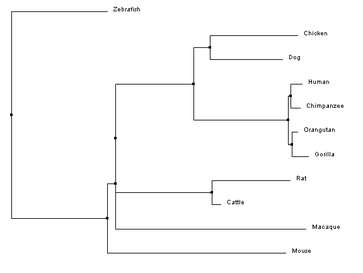
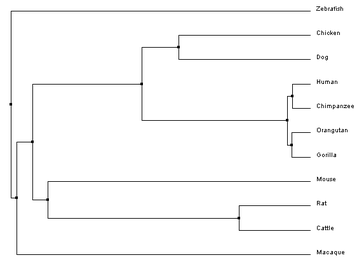
Clustal Omega requires FASTA formatted sequences to be entered for each gene of interest. These FASTA sequences for the FANCL homologs were obtained from the NCIB gene database and are compiled in the document below. Upon completion of the alignment, the program provides a complete alignment of the gene sequence that can be modified to highlight multiple aspects of the similarities and differences at each nucleotide position in the sequence. Below is the sequence alignment for the human FANCL protein and the nine homologous genes. Constructing phylogenetic treesAfter aligning sequences, phylogenetic trees can be constructed to predict the relatedness of each gene to the others. Clustal Omega calculates these phylogenetic trees directly within the program used to align the sequences, and can predict trees using a percent identity sequence similarity scoring method in combination with two different tree construction models.
Analysis of ResultsThe results of the phylogenetics analysis were not what would be expected of a typical tree drawing. The selection of organisms was thought out to provide multiple primates, which would provide pretty similar sequences, and then other species from mammals, to birds, to fish, to increase the evolutionary divergence and potentially increase the sequence disparity. However, the purpose of the selection of organisms did not correlate to the expected results. Of note is the rhesus macaque, which was assumed to be a similar sequence to the other primates, but was outside of even the chicken. Although it is difficult to extrapolate why this is from nucleotide sequence analysis, looking at protein phylogeny, a clearer picture of the difference in the macaque sequence may come to light.
|
References
[1] Sievers F, Wilm A, Dineen DG, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, Thompson JD, Higgins DMolecular Systems Biology 7 Article number: 539 doi:10.1038/msb.2011.75
[2] Barton, N. H., Briggs, D. E., Eisen, J. A., Goldstein, D. B., & Patel, N. H. (2007). Evolution. N.p.: Cold Spring Harbor.
[2] Barton, N. H., Briggs, D. E., Eisen, J. A., Goldstein, D. B., & Patel, N. H. (2007). Evolution. N.p.: Cold Spring Harbor.
File Resources
| fancl_gene.txt | |
| File Size: | 19 kb |
| File Type: | txt |